16S sequencing protocol
Purpose & Assumptions
The purpose of this guide is to describe the primary differences in analyzing sequences generated using the Earth Microbiome Project (EMP) or standard Illumina protocols. In other words, this post will describe the practical aspects and applications with respect to analyzing the data, rather than the theory behind the process, study design, etc.
At the Center for Quantitative Life Sciences (CQLS) at OSU, we explicitly support two 16S sequencing protocols. Here are relevant links for both processes:
16S Primer Targets
Different 16S sequencing protocols target different hypervariable regions of the 16S rRNA gene sequence. The EMP protocol targets the V4 region, while the Illumina protocol targets the V3-V4 region.
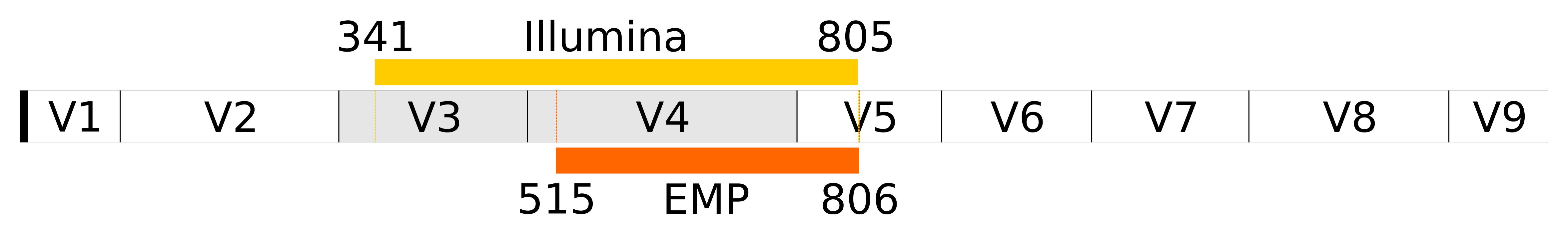
The Illumina 16S protocol is based on the primers from Klindworth et al. 2012, while the EMP protocol has been revised several times, currently based on the 515F (Parada)–806R (Apprill), forward-barcoded primer set (links to these publications can be found on the EMP page).
The Illumina 16S protocol leverages the same sequences as the Nextera XT library prep process used for sequencing genomic data. However, the sequences are attached to the inserts using a two-step PCR process, rather than the standard tagmentation process. The Illumina 16S products utilize a dual index barcode process.
Conversely, the EMP 16S protocol uses no Nextera XT sequences in the process, but rather only leverages the P5 and P7 sequences at the 5' and 3' ends, respectively, of the PCR product, such that the PCR amplicons can be attached to a Illumina flowcell and then sequenced. The only barcodes are found on the forward primers (single index).
Data processing
The question becomes, how does one process the data generated using these two protocols?
The DADA2 tutorial is relatively extensive and is a good guide for analyzing 16S sequences.
Here is an excerpt from the DADA2 tutorial as it relates to the differences in these two sequencing processes:
Starting point
This workflow assumes that your sequencing data meets certain criteria:
- Samples have been demultiplexed, i.e. split into individual per-sample fastq files.
- Non-biological nucleotides have been removed, e.g. primers, adapters, linkers, etc.
- If paired-end sequencing data, the forward and reverse fastq files contain reads in matched order.
If these criteria are not true for your data (are you sure there aren’t any primers hanging around?) you need to remedy those issues before beginning this workflow. See the FAQ for recommendations for some common issues.
The CQLS, by default, will demultiplex all multiplexed samples using the barcodes as provided when samples were submitted for library prep and/or sequencing. Reads will be in matched order after this process. The final remaining item, the presence of primers, adapters, etc. will depend on which protocol was used for sequencing.
If primers (especially those with degeneracy) are present, those must be removed for several reasons, including:
- Mismatches between the primer and 16S target can occur, introducing additional sequence variance in the output
- Additional sequence variance can cause difficulty in terms of DADA2 calculating the expected error rates in the reads
In order to determine if you have primers in your output, it is helpful to understand the differences between how the reads are generated in the two 16S sequencing protocols described here.
The CQLS does not routinely remove PCR primer sequences as part of the standard sequencing process. Feel free to contact me if you do have questions regarding primer removal, especially after reading this post.
Despite this fact, you may find that you have no primer sequences in your EMP protocol raw output.
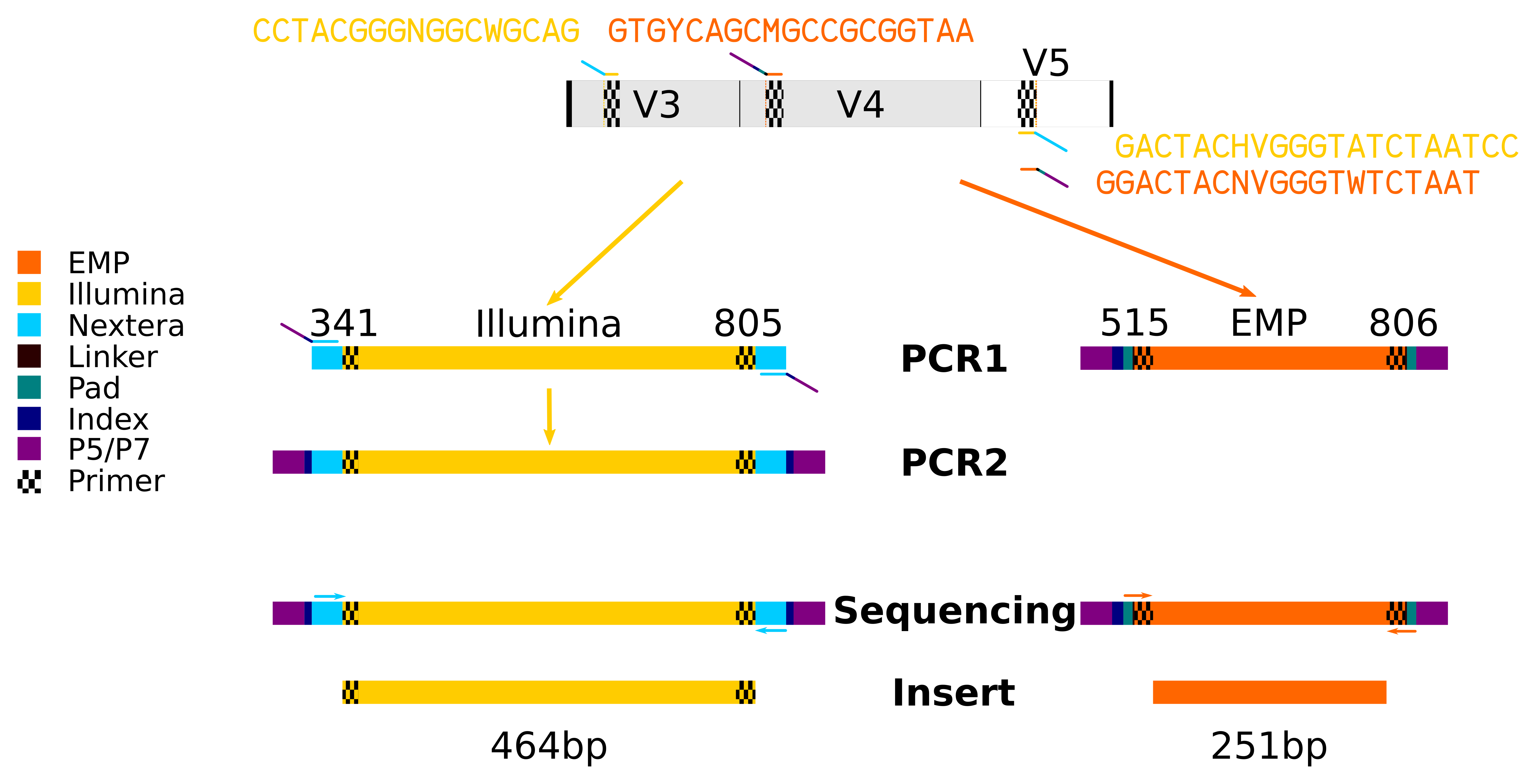
Put simply, the most important determinant of primer sequences in the raw output from the Illumina MiSeq sequencer is the type and target of the sequencing primer(s) that is loaded onto the sequencer. In the Illumina protocol, the sequencing primers used target the Nextera sequence, upstream of the 16S primers in the inserts. Conversely, the EMP protocol has sequencing primers that anneal to the 16S primer sequences and upstream linker/pad sequences. Therefore, the 5' end of the raw reads in the Illumina protocol contains the 16S primer sequences, while the EMP protocol raw reads omit those primer sequences.
Note: The EMP page describes the sequencing primers and their inclusion in the sequencing protocol.
You’ll see in the sequencing diagram that there are several differences between the two 16S sequencing library preparation protocols. The similarities/differences are described in a table below:
| Protocol | Illumina | EMP |
|---|---|---|
| Target | V3-V4 | V4 |
| Target Length (bp) | 464 | 251 |
| Forward Primer | CCTACGGGNGGCWGCAG | GTGYCAGCMGCCGCGGTAA |
| Forward Primer Length (bp) | 17 | 19 |
| Reverse Primer | GACTACHVGGGTATCTAATCC | GGACTACNVGGGTWTCTAAT |
| Reverse Primer Length (bp) | 21 | 20 |
| # PCR Steps | 2 | 1 |
| Flowcell Sequencing Target | Nextera Sequence | 16S Primer |
| 16S Primer in Output | Yes | No |
Using the filterAndTrim() function of DADA2
The filterAndTrim() function has many settings that need to be manually set by
the user to produce the highest quality analysis.
Primer sequences can be removed using multiple techniques, but when using DADA2,
the most straightfoward method is during the filterAndTrim() step of the
pipeline. The trimL option can be used to systematically trim off the primer
sequences at the 5' end of the reads. For the Illumina protocol, one can set
trimL = c(17,21) to trim off the primers as found in a typical Illumina 16S
sequencing run. As stated above, no trimL needs to be supplied during the EMP
analysis.
Additionally, one can examine the FASTQC (or aggregated multiqc) output to find
the most appropriate truncation length (truncLen) setting. In general, one can
expect to set truncLen = c(240, 200) for the EMP protocol. For the Illumina
protocol, one must leave a longer sequence in order to have enough overlap to
merge the reads. Illumina suggests at least 50bp of overlap, while the DADA2
authors expect at least 20bp of overlap. (464+50)/2 reads would mean an expected
length of 257bp for each the forward and reverse reads for the Illumina
protocol. At minimum, one will not be able to trim to the 240, 200 as the FASTQ
quality scores suggest. I generally set truncLen = c(250, 250) for the Illumina
protocol, which is a bit shorter than suggested but still works most of the time.
You can experiment with leaving longer truncLen settings and see which works
best for your particular dataset.
Identifying primers using the command line
Let’s say you have sequences and you aren’t sure if you have primers in your reads.
I wrote a small script that can be used as a reference for peeking at the 5' end of the reads:
|
|
Example usage: fastqPeek R1.fastq.gz 5 25 4000 Would print the first 1000
(4000/4) sequences on the 5' end and pull out the first 25 bases of the seqs.
Your primers should be in the first 20 bases or so of most sequencing protocols.
You can manually run the commands starting with the zcat or cat on your own
as well. One idea would be to change the pipeline at the sort -u command, e.g.
zcat $infile | head -n $lines | sed -n '2~4p' | cut -c $pos | sort | uniq -c
Which would count the number of times each sequence is seen. If you see a large number for a particular set of seqs, that either means there is a high abundance sequence in the first n number of lines in your file, or you have a primer seq still in your reads.
References: